day7 GEO下载多样本数据的聚类注释
原创R基础知识补充:
字符串处理函数 - str_split_i() - str_sub() - str_remove() - str_detect() - str_replace() 文件处理函数 - dir(pattern = ".R$") - file.create(" ") - file.exists(" ") - file.remove(" ") - f = paste0("douhua",1:10,".txt"); file.create(f) - lappy函数 自定义函数 my_install = function(pkg){ if (!require(pkg,character.only = T))install.packages(pkg,update = F,ask = F) }
my_install("tidyr") my_install("dplyr") my_install("stringr") ps = c("tidyr","dplyr","stringr") lapply(ps, my_install) #利用好lappy写简洁函数;有点>for循环的意思?
ifelse 函数 :根据逻辑值是T还是F产生不同的值
数据获取+文件名修改
数据来自GEO的GSE231920,有3个treat,3个control样本
- 全部下载并解压
untar("GSE231920_RAW.tar",exdir = "GSE231920_RAW") #解包- 改名 利用lapply套自定义函数实现了批量操作
- 为每个样本创建单独的文件夹
- 把每个样本的三个文件复制进去
- 所有文件改名,去掉前缀
library(stringr)
fs = paste0("GSE231920_RAW/",dir("GSE231920_RAW/"));fs #所有文件的路径
samples = dir("GSE231920_RAW/") %>% str_split_i(pattern = "_",i = 2) %>% unique();samples #str_split_i()进行拆分,并选择想要的第2个部分(i=2),即sample
##############################
#01自定义函数批量创建文件目录 #s即sample
lapply(samples, function(s){
sdir = paste0("02_data/",s)
if(!file.exists(sdir))dir.create(sdir,recursive = T)
})
#02自定义函数批量移动文件 #匹配到相同的sample名称就copy
lapply(fs, function(s){
for(i in 1:length(samples)){
if(str_detect(s,samples[[i]])){ file.copy(s,paste0("02_data/",samples[[i]])) }
}
})
#03去掉文件的前缀 #修改文件名时,路径时要从工作目录之下开始写的
on = paste0("01_data/",dir("01_data/",recursive = T));on
nn = str_remove(on,"GSM\\d+_sample\\d_");nn
file.rename(on,nn)
##############################批量读取和抽样保存
- 批量读取
接下来就是批量读取文件,前面我们知道利用
Read10X()函数读取三个固定文件的目录名称,利用for循环每读取一个samplepda,就创建一个对象SeuratObjectsce,并存储在列表中scelist:
f = dir("01_data/")
scelist = list() #创建空的列表,下面的for循环每执行一次,scelist里面就会多一个元素。
for(i in 1:length(f)){
pda <- Read10X(paste0("01_data/",f[[i]]))
scelist[[i]] <- CreateSeuratObject(counts = pda,
project = f[[i]],
min.cells = 3,
min.features = 200)
print(dim(scelist[[i]]))
}- 合并 然后我们再把列表中的每个sce合并,合并后每个样本的表达矩阵是一个独立的的layer,JoinLayers是合并为一个表达矩阵:
sce.all = merge(scelist[[1]],scelist[-1]) #合并多个对象
sce.all = JoinLayers(sce.all) - 抽样保存
set.seed(335)
sce.all = subset(sce.all,downsample=700)
save(sce.all,file = rdaf)质控+降维+注释
- 质控
#除了线粒体基因,核糖体和红细胞基因也是常见的过滤指标
sce.all[["percent.mt"]] <- PercentageFeatureSet(sce.all, pattern = "^MT-")
sce.all[["percent.rp"]] <- PercentageFeatureSet(sce.all, pattern = "^RP[SL]")
sce.all[["percent.hb"]] <- PercentageFeatureSet(sce.all, pattern = "^HB[^(P)]")
VlnPlot(sce.all,
features = c("nFeature_RNA",
"nCount_RNA",
"percent.mt",
"percent.rp",
"percent.hb"),
ncol = 3,pt.size = 0.5, group.by = "orig.ident")
#根据小提琴图指定指标去掉离群值
sce.all = subset(sce.all,percent.mt < 20&
nCount_RNA < 40000 &
nFeature_RNA < 6000)
table(sce.all@meta.data$orig.ident)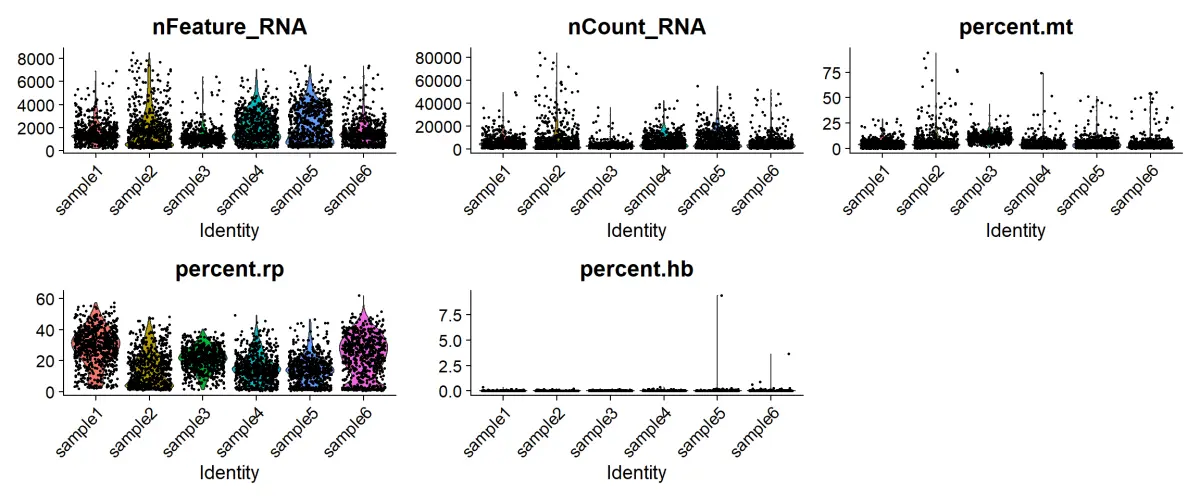
- 降维聚类,多样本使用harmony,它需要的计算资源少且准确程度高,是最受欢迎的方法
f = "obj.Rdata"
library(harmony)
if(!file.exists(f)){
sce.all = sce.all %>%
NormalizeData() %>%
FindVariableFeatures() %>%
ScaleData(features = rownames(.)) %>%
RunPCA(pc.genes = VariableFeatures(.)) %>%
RunHarmony("orig.ident") %>%
FindNeighbors(dims = 1:15, reduction = "harmony") %>%
FindClusters(resolution = 0.5) %>%
RunUMAP(dims = 1:15,reduction = "harmony") %>%
RunTSNE(dims = 1:15,reduction = "harmony")
save(sce.all,file = f)
}
load(f)
ElbowPlot(sce.all)
p1 = DimPlot(sce.all, reduction = "umap",label = T,pt.size = 0.5)+ NoLegend();p1
DimPlot(sce.all, reduction = "umap",pt.size = 0.5,group.by = "orig.ident") #按照样本来分配颜色。可以看看去除样本间批次效应的效果如何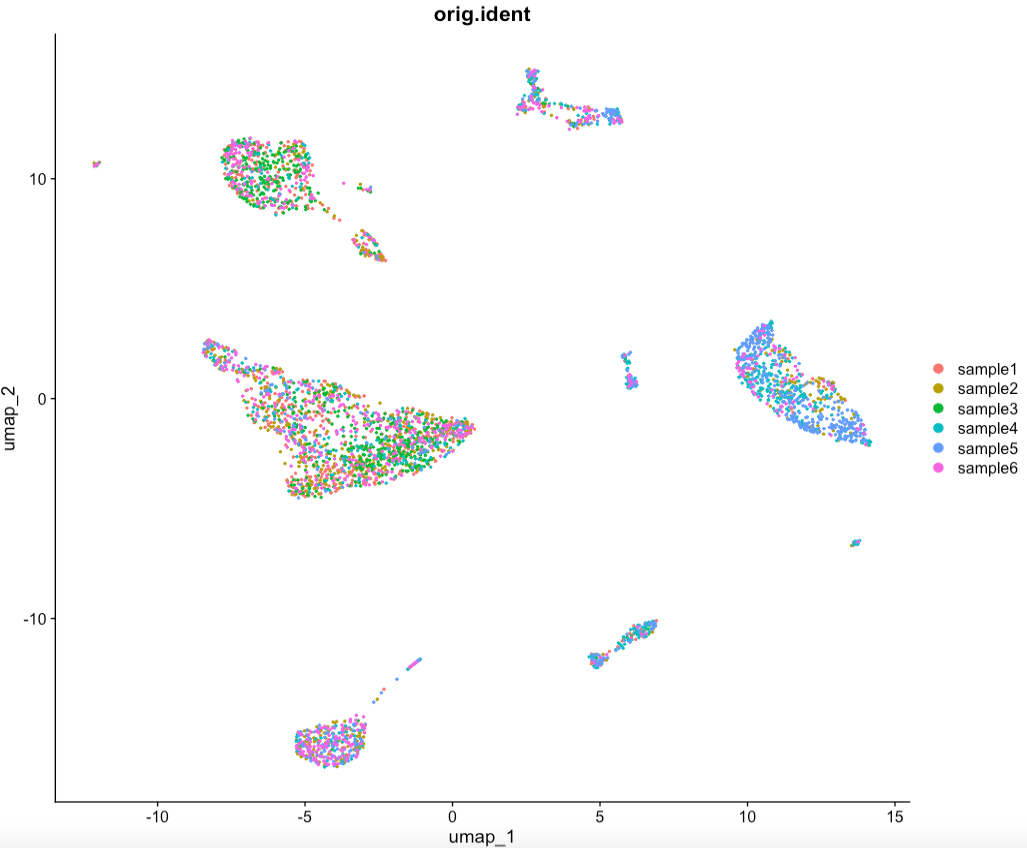
- 注释
library(celldex)
library(SingleR)
f = "../day5-6/ref_BlueprintEncode.RData"
if(!file.exists(f)){
ref <- celldex::BlueprintEncodeData()
save(ref,file = f)
}
ref <- get(load(f))
library(BiocParallel)
scRNA = sce.all
test = scRNA@assays$RNA$data
pred.scRNA <- SingleR(test = test,
ref = ref,
labels = ref$label.main,
clusters = scRNA@active.ident)
pred.scRNA$pruned.labels
new.cluster.ids <- pred.scRNA$pruned.labels
names(new.cluster.ids) <- levels(scRNA)
scRNA <- RenameIdents(scRNA,new.cluster.ids)
save(scRNA,file = "scRNA.Rdata")
p2 <- DimPlot(scRNA, reduction = "umap",label = T,pt.size = 0.5) + NoLegend()
p1+p2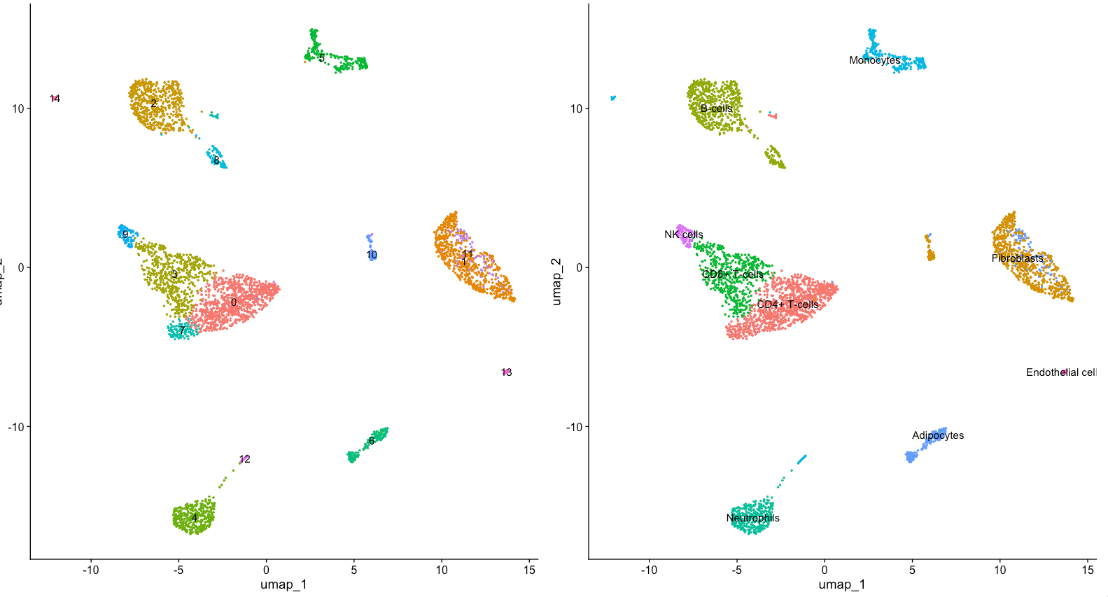
分组可视化和组间细胞比例比较
- 添加分组信息 library(tinyarray) edat = geo_download("GSE231920") pd = edat$pd 可以自行提取GSE的分组信息
scRNA$seurat_annotations = Idents(scRNA) #加入细胞类型列
#利用ifelse函数判断是处理组还是对照组
scRNA$group = ifelse(scRNA$orig.ident %in% c("sample1","sample2","sample3"), "treat","control")
DimPlot(scRNA, reduction = "umap", group.by = "group")- 计算每个亚群的细胞数量占全部细胞的比例
cell_counts <- table(Idents(scRNA))
cell.all <- cbind(cell_counts = cell_counts,
cell_Freq = round(prop.table(cell_counts)*100,2))
#各组中每种细胞的数量和比例
cell.num.group <- table(Idents(scRNA), scRNA$group)
cell.freq.group <- round(prop.table(cell.num.group, margin = 2) *100,2)
cell.all = cbind(cell.all,cell.num.group,cell.freq.group)
cell.all = cell.all[,c(1,3,4,2,5,6)]
colnames(cell.all) = paste(rep(c("all","control","treat"),times = 2),
rep(c("count","freq"),each = 3),sep = "_")
cell.all差异分析
- 组间差异基因
以NK细胞为例,
FindMarkers()找差异基因dfmarkers
sub.obj = subset(scRNA,idents = "NK cells")
dfmarkers <- FindMarkers(scRNA, ident.1 = "treat", group.by = "group",min.pct = 0.25, logfc.threshold = 0.25,verbose = F)
head(dfmarkers) - 保守 marker基因,
FindConservedMarkers()找NK细胞与其他亚群细胞的相比的差异保守基因sub.markers
if(!require("multtest"))BiocManager::install('multtest')
if(!require("metap"))install.packages('metap')
sub.markers <- FindConservedMarkers(scRNA, ident.1 = "NK cells", grouping.var = "group", min.pct = 0.25, logfc.threshold = 0.25,verbose = F)
head(sub.markers)- 组间比较气泡图,把感兴趣的基因进行
markers.to.plot = c("CD3D", "CREM", "HSPH1", "SELL", "GIMAP5", "CACYBP", "GNLY", "NKG7", "CCL5",
"CD8A", "MS4A1", "CD79A", "MIR155HG", "NME1", "FCGR3A", "VMO1", "CCL2", "S100A9", "HLA-DQA1",
"GPR183", "PPBP", "GNG11", "HBA2", "HBB", "TSPAN13", "IL3RA", "PRSS57")
DotPlot(scRNA, features = markers.to.plot, cols = c("blue", "red"),
dot.scale = 8, split.by = "group") +
RotatedAxis()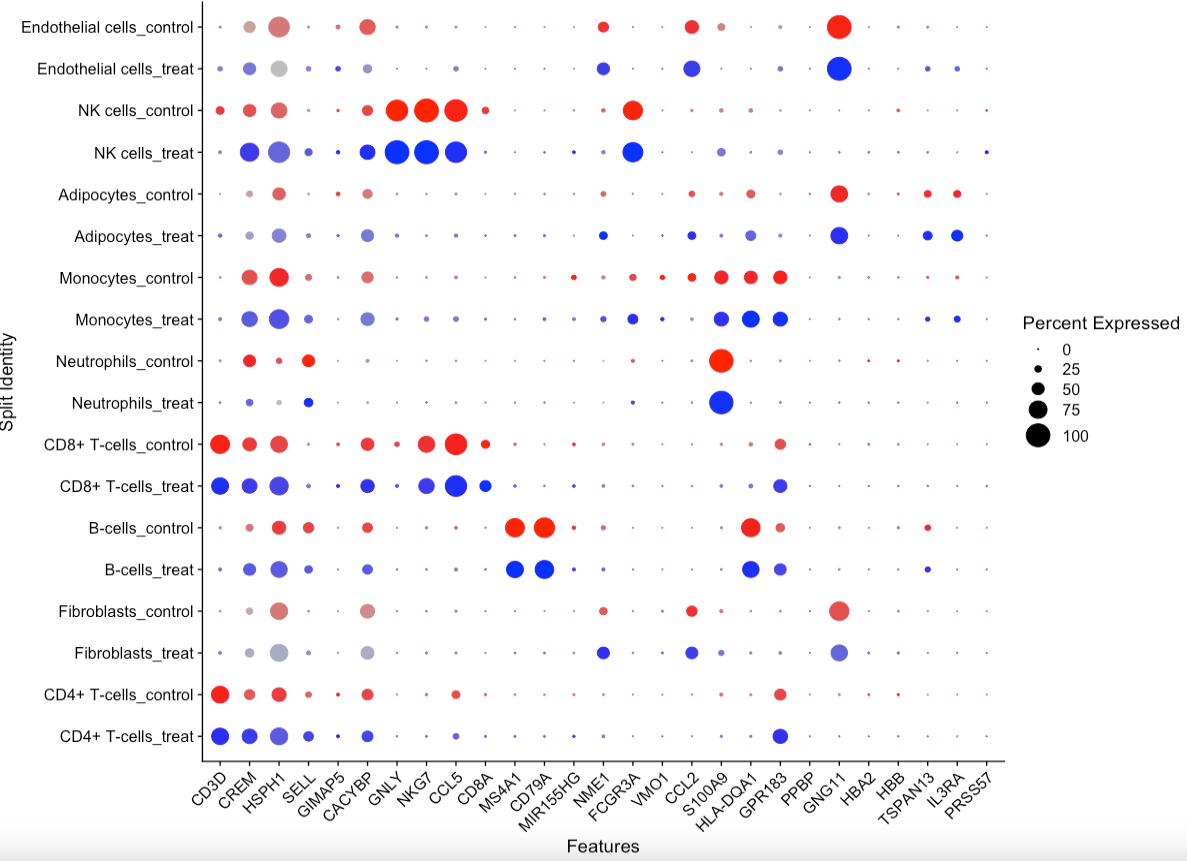
- 回溯细胞图和小提琴图
#FeaturePlot
FeaturePlot(scRNA, features = c("CD3D", "GNLY", "IFI6"), split.by = "group", max.cutoff = 3, cols = c("grey","red"), reduction = "umap")
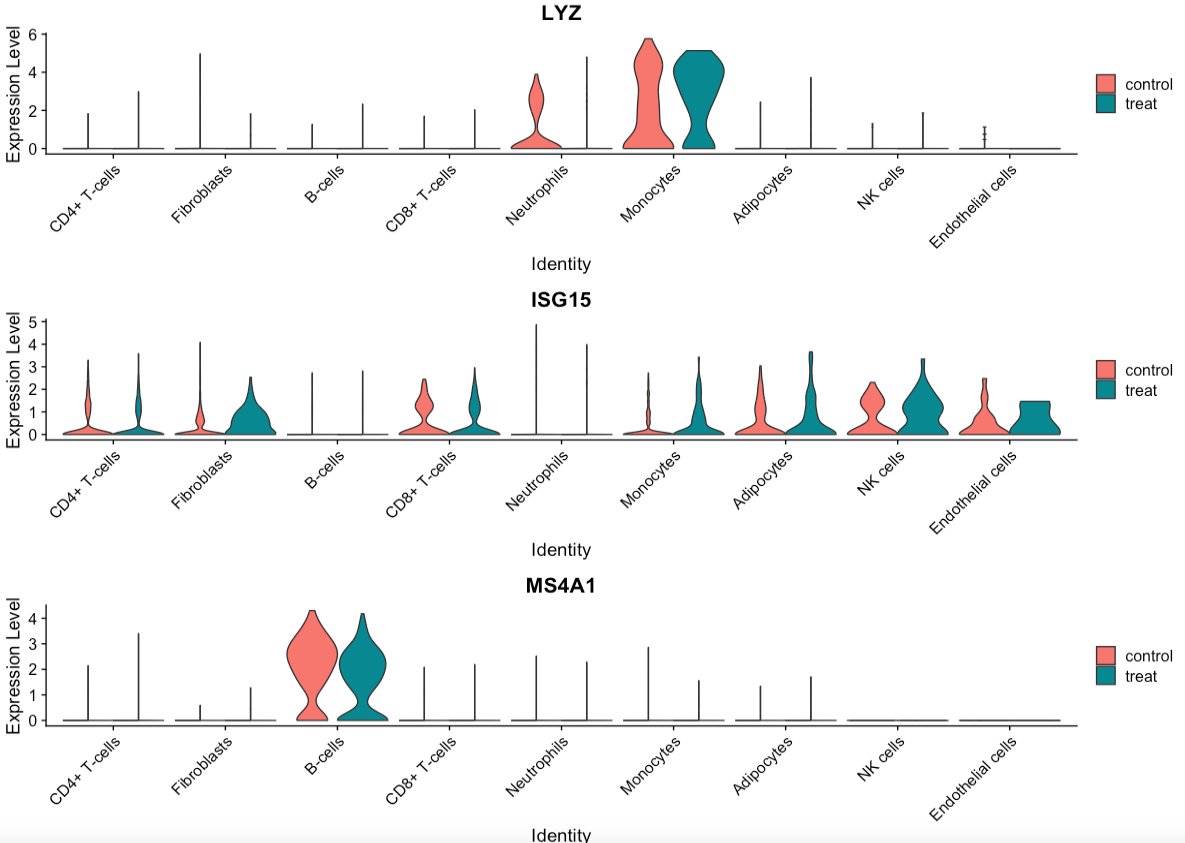
#小提琴图
plots <- VlnPlot(scRNA, features = c("LYZ", "ISG15", "MS4A1"), split.by = "group", group.by = "seurat_annotations",
pt.size = 0, combine = FALSE)
library(patchwork)
wrap_plots(plots = plots, ncol = 1)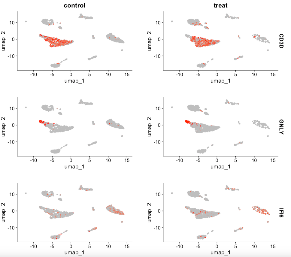
伪bulk转录组分析
把单细胞数据整合为常规转录组数据的方式
#每个组需要多个样本才能做,如两组各有3个
bulk <- AggregateExpression(scRNA, return.seurat = T, slot = "counts", assays = "RNA", group.by = c("seurat_annotations","orig.ident", "group"))
colnames(bulk)#整合成了多个“样本”
sub <- subset(bulk, seurat_annotations == "CD8+ T-cells")
colnames(sub)
Idents(sub) <- "group"
de_markers <- FindMarkers(sub, ident.1 = "treat", ident.2 = "control", slot = "counts", test.use = "DESeq2",
verbose = F)
de_markers$gene <- rownames(de_markers)
k1 = de_markers$avg_log2FC< -1 & de_markers$p_val <0.01
k2 = de_markers$avg_log2FC> 1 & de_markers$p_val <0.01
de_markers$change <- ifelse(k1,"down",ifelse(k2,"up","not"))de_markers火山图可视化
library(ggplot2)
ggplot(de_markers, aes(avg_log2FC, -log10(p_val),color = change)) +
geom_point(size = 2, alpha = 0.5) +
geom_vline(xintercept = c(1,-1),linetype = 4)+
geom_hline(yintercept = -log10(0.01),linetype = 4)+
scale_color_manual(values = c("#2874C5", "grey", "#f87669"))+
theme_bw() +
ylab("-log10(unadjusted p-value)") 原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
评论
作者已关闭评论
推荐阅读
目录


