decoupleR:一站式scRNA通路活性分析及转录因子活性推断
decoupleR:一站式scRNA通路活性分析及转录因子活性推断

KS科研分享与服务-TS的美梦
发布于 2026-04-29 13:44:48
发布于 2026-04-29 13:44:48
decoupleR是提供了一套统一框架下的多种计算方法,用于从组学数据中推断和提取生物学活性的R包。能够进行转录组数据通路活性分析以及转录因子活性分析。从通路活性分析角度来看,是一个集合多种方法的框架,比如常见的基因集评分方法它都具备;从转录因子活性分析来看,是SCENIC的减配版平替。decoupleR依托先验知识数据库,分析比较简单。
markdown版详细内容已发布微信VIP群,请自行下载!
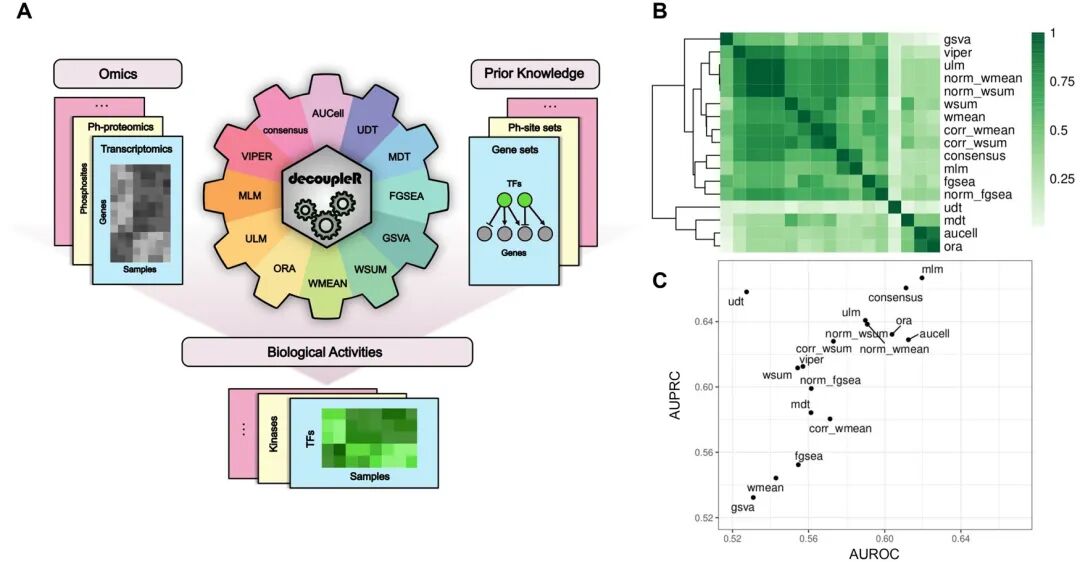
decoupleR workflow
Github:https://github.com/saezlab/decoupleR/?tab=readme-ov-file
Citation:Badia-i-Mompel P., Vélez Santiago J., Braunger J., Geiss C., Dimitrov D., Müller-Dott S., Taus P., Dugourd A., Holland C.H., Ramirez Flores R.O. and Saez-Rodriguez J. 2022. decoupleR: ensemble of computational methods to infer biological activities from omics data. Bioinformatics Advances. https://doi.org/10.1093/bioadv/vbac016
安装包:
#install.packages('remotes')
#remotes::install_github('saezlab/decoupleR')
library(decoupleR)
packageVersion("decoupleR")setwd("/home/tq_ziv/data_analysis/R版decoupleR")
library(decoupleR)
library(dplyr)
library(tibble)
library(tidyr)
library(ggplot2)
library(pheatmap)
library(ggrepel)
library(Seurat)查看下decoupleR包所有的method:
## # A tibble: 12 × 2
## Function Name
## <chr> <chr>
## 1 run_aucell AUCell
## 2 run_consensus Consensus score between methods
## 3 run_fgsea Fast Gene Set Enrichment Analysis (FGSEA)
## 4 run_gsva Gene Set Variation Analysis (GSVA)
## 5 run_mdt Multivariate Decision Trees (MDT)
## 6 run_mlm Multivariate Linear Model (MLM)
## 7 run_ora Over Representation Analysis (ORA)
## 8 run_udt Univariate Decision Tree (UDT)
## 9 run_ulm Univariate Linear Model (ULM)
## 10 run_viper Virtual Inference of Protein-activity by Enriched Regulon anal…
## 11 run_wmean Weighted Mean (WMEAN)
## 12 run_wsum Weighted Sum (WSUM)加载单细胞数据:
load("~/data_analysis/R版decoupleR/sce_cca_cellltype.RData")DimPlot(sce_cca,split.by = 'orig.ident')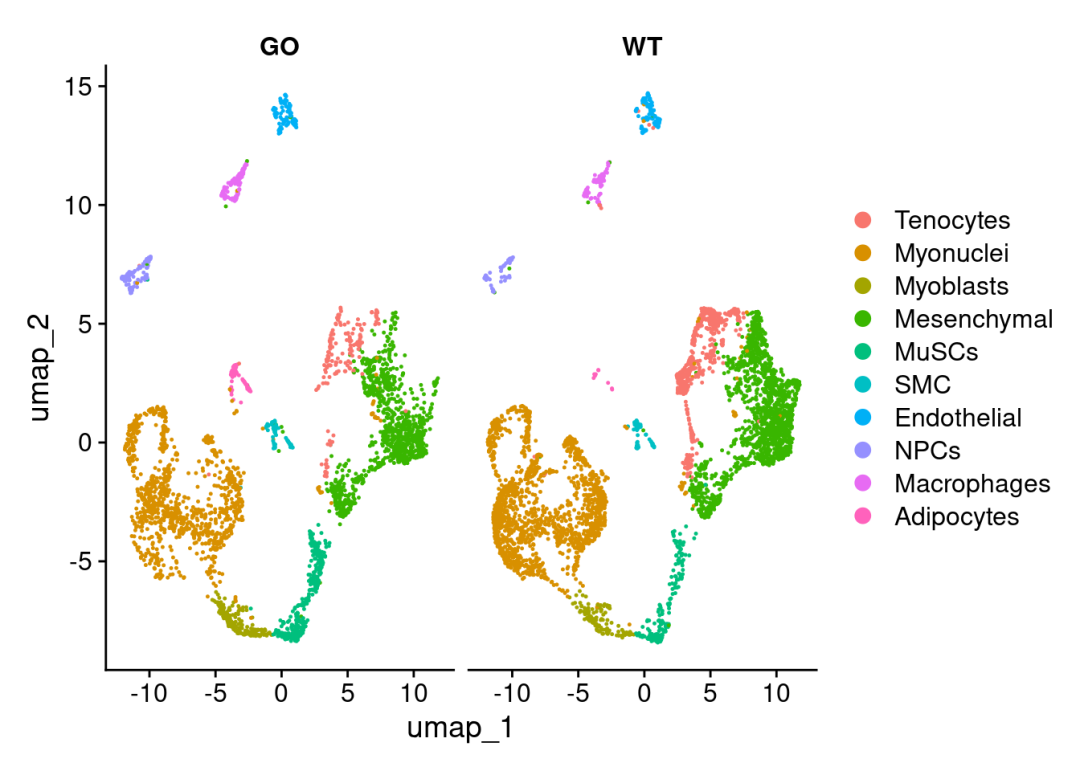
分析1-scRNA通路活性推断
首先获取PROGENyhttps://saezlab.github.io/progeny/通路网络,PROGENy: Pathway RespOnsive GENes for activity inference,是利用大量公开可用的信号通路干扰实验汇编而成的资料,为人类和小鼠生成了通路响应基因的共同核心资源。这些基因与任何统计方法相结合,可用于从总体或单细胞转录组学中推断通路活性。也就是说,PROGENy是基于先验知识的总结,是一个综合资源库,其中包含经过整理的通路及其目标基因,以及每个相互作用的权重。在decoupleR中,可以使用get_progeny()函数获取资源数据,用于下游的分析推断。获取的内容是一个dataframe,包括source pthway,target gene,权重及P值。get_progeny()获取的通路包括14种pathway,都是常见熟悉的通路,简介如下:
Androgen: involved in the growth and development of the male reproductive organs. EGFR: regulates growth, survival, migration, apoptosis, proliferation, and differentiation in mammalian cells. Estrogen: promotes the growth and development of the female reproductive organs. Hypoxia: promotes angiogenesis and metabolic reprogramming when O2 levels are low. JAK-STAT: involved in immunity, cell division, cell death, and tumor formation. MAPK: integrates external signals and promotes cell growth and proliferation. NFkB: regulates immune response, cytokine production and cell survival. p53: regulates cell cycle, apoptosis, DNA repair and tumor suppression. PI3K: promotes growth and proliferation. TGFb: involved in development, homeostasis, and repair of most tissues. TNFa: mediates haematopoiesis, immune surveillance, tumour regression and protection from infection. Trail: induces apoptosis. VEGF: mediates angiogenesis, vascular permeability, and cell migration. WNT: regulates organ morphogenesis during development and tissue repair.
很显然,内置的PROGENy并不能满足个性化的分析需求,比如自己关注的通路不在,那么还能使用decoupleR进行通路活性推断吗?答案是肯定的,自定义数据只需要按照decoupleR需求的格式整理数据即可。
net <- decoupleR::get_progeny(organism = 'mouse', #只支持“human”或者“mouse”
top = 500) #返回每条通路top n的基因通路活性推断:
单细胞数据需要标准化后的表达矩阵:推断矩阵中每个cells的通路活性评分。
mat <- GetAssayData(sce_cca, layer = 'data')decoupleR的方法基本分为两类,一类是线性模型,例如run_ulm,run_mlm,run_mdt,run_viper,run_udt,这些都利用“regulator–target网络+基因表达”,推断上游调控因子活性。另一类是基因集富集 打分型,例如run_aucell,run_gsva,run_fgsea,run_ora等,这些更像“传统基因集富集”。run_consensus是一种组合方法,输入多个方法的结果,对它们的得分/排序做融合得到更稳健的总体结果。具体的方法介绍参考:
https://academic.oup.com/bioinformaticsadvances/article/2/1/vbac016/6544613?login=false.这里使用Multivariate Linear Model (mlm) 方法推断通路富集分数,对于数据集中的每个样本,它拟合一个线性模型,该模型基于所有通路的通路 - 基因相互作用权重来预测观察到的基因表达。一旦拟合完成,所获得的斜率的t值即为得分。如果该值为正,则解释为该通路处于活跃状态;如果为负,则解释为该通路处于不活跃状态。
# Run mlm
acts <- decoupleR::run_mlm(mat = mat,
net = net,
.source = 'source', #输入net中的source列
.target = 'target', #输入net中的target列
.mor = 'weight', #输入net中的权重列
minsize = 5)将结果返回seurat:分析得到的acts是一个长数据,将其转化为宽数据矩阵,添加到seurat中新的assay。
sce_cca[['pathwaysmlm']] <- acts %>%
tidyr::pivot_wider(id_cols = 'source',
names_from = 'condition',
values_from = 'score') %>%
tibble::column_to_rownames(var = 'source') %>%
Seurat::CreateAssayObject(.)DefaultAssay(sce_cca) <- 'pathwaysmlm'
# Scale the data
sce_cca <- Seurat::ScaleData(sce_cca)sce_cca@assays$pathwaysmlm@data <- sce_cca@assays$pathwaysmlm@scale.datacolors <- rev(RColorBrewer::brewer.pal(n = 11, name = "RdBu")[c(2, 10)])
FeaturePlot(sce_cca, features = c("Trail")) +
ggplot2::scale_colour_gradient2(low = colors[1], mid = 'white', high = colors[2]) +
ggplot2::ggtitle('Trail activity')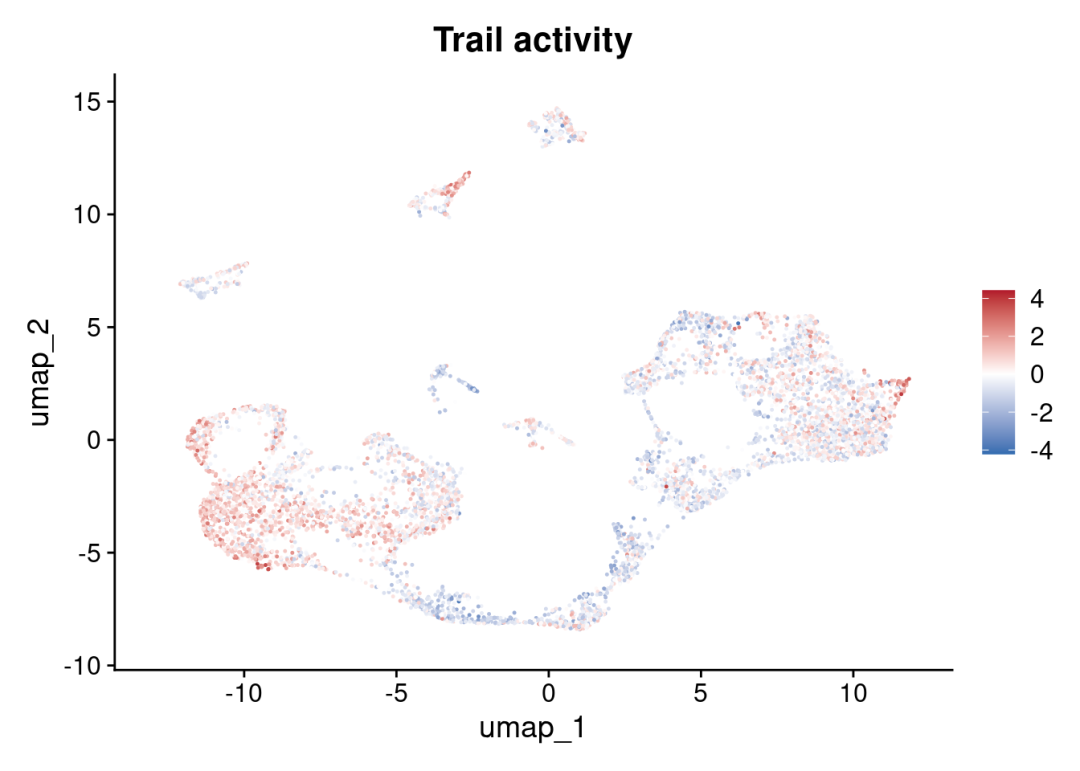
colors <- rev(RColorBrewer::brewer.pal(n = 11, name = "RdBu")[c(2, 10)])
FeaturePlot(sce_cca, features = c("Trail"),split.by = 'orig.ident') &
ggplot2::scale_colour_gradient2(low = colors[1], mid = 'white', high = colors[2])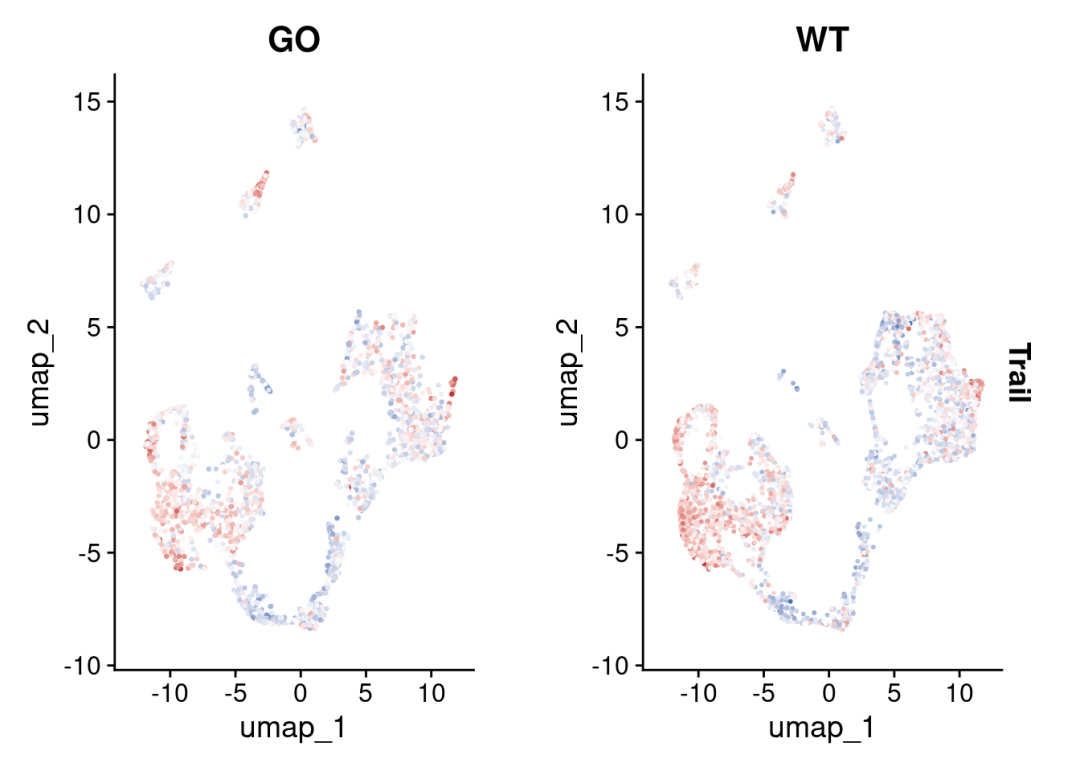
#例如只看一个细胞类型
Idents(sce_cca) <- "celltype"
MUscs <- subset(sce_cca, idents = "MuSCs")
#看看celltype内通路活性差异
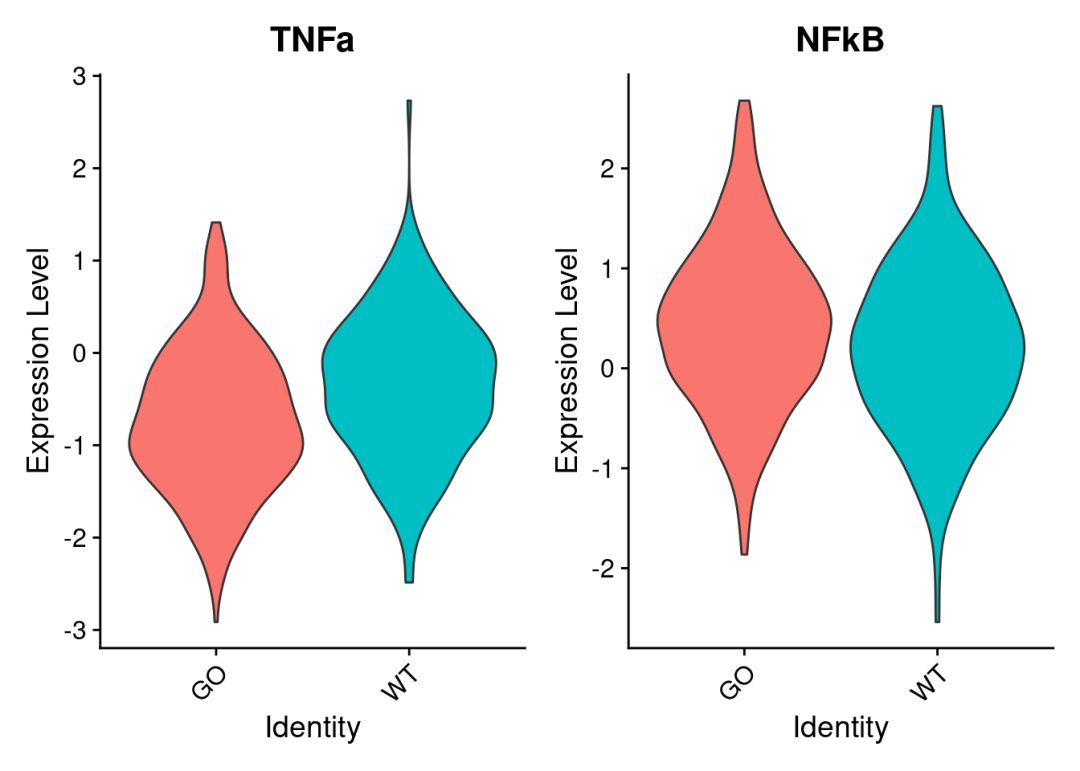
VlnPlot(
MUscs,
features = c("TNFa", "NFkB"),
group.by = "orig.ident",
pt.size = 0
)
sce_cca$group_cells <- paste0(sce_cca$orig.ident,"_",sce_cca$celltype)
Idents(sce_cca) <- "group_cells"
# Extract activities from object as a long dataframe
df <- t(as.matrix(sce_cca@assays$pathwaysmlm@data)) %>%
as.data.frame() %>%
dplyr::mutate(cluster = Seurat::Idents(sce_cca)) %>%
tidyr::pivot_longer(cols = -cluster,
names_to = "source",
values_to = "score") %>%
dplyr::group_by(cluster, source) %>%
dplyr::summarise(mean = mean(score))
# Transform to wide matrix
top_acts_mat <- df %>%
tidyr::pivot_wider(id_cols = 'cluster',
names_from = 'source',
values_from = 'mean') %>%
tibble::column_to_rownames(var = 'cluster') %>%
as.matrix()
# Color scale
colors <- rev(RColorBrewer::brewer.pal(n = 11, name = "RdBu"))
colors.use <- grDevices::colorRampPalette(colors = colors)(100)
my_breaks <- c(seq(-1.25, 0, length.out = ceiling(100 / 2) + 1),
seq(0.05, 1.25, length.out = floor(100 / 2)))
top_acts_mat <- top_acts_mat[c("WT_Tenocytes","GO_Tenocytes",
"WT_Myonuclei","GO_Myonuclei",
"WT_Myoblasts","GO_Myoblasts",
"WT_Mesenchymal","GO_Mesenchymal",
"WT_MuSCs","GO_SMC",
"WT_SMC","GO_SMC",
"WT_Endothelial","GO_Endothelial",
"WT_NPCs","GO_NPCs",
"WT_Macrophages","GO_Macrophages",
"WT_Adipocytes","GO_Adipocytes"),]
pheatmap::pheatmap(mat = top_acts_mat,
color = colors.use,
border_color = "white",
breaks = my_breaks,
cellwidth = 20,
cellheight = 20,
treeheight_row = 20,
treeheight_col = 20,
cluster_rows = F) 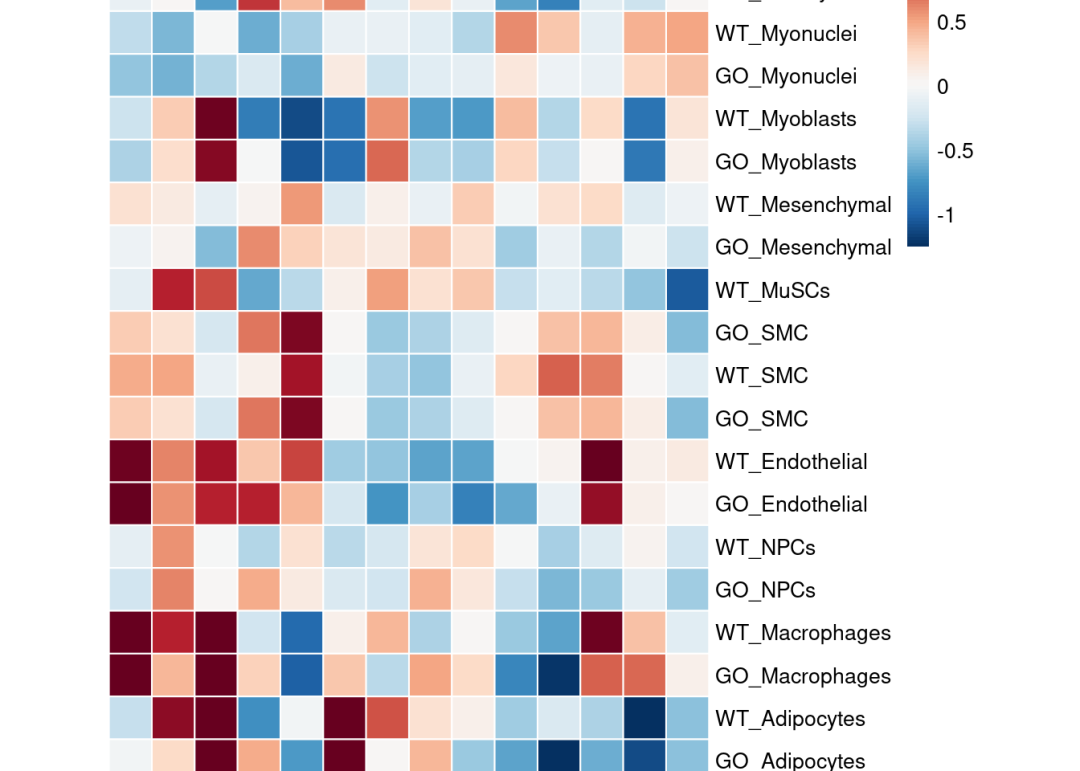
分析2-scRNA转录因子活性推断
转录因子通过调控下游靶基因的转录,在细胞命运决定和疾病发生发展中发挥核心作用。从转录组数据可靠地推断TF活性,成为解析细胞异质性和识别关键调控因子的关键问题。我们介绍最多,使用非常多的方法SCENIC(pySCENIC)是有效的方法,但是流程复杂,计算工作量大,且对于普通bulk RNA数据效果不好(样本量不足)。decoupleR利用利用现成网络+多算法打分,进行单细胞及BULK转录组数据的转录因子活性推断。数据库来源于先验知识,使用CollecTRI network,其中包含从12个不同来源精心整理的转录因子及其转录靶点的集合,是DoRothEA(DoRothEA: collection of human and mouse regulons) 网络的拓展,但是与其和其他基于文献的基因调控网络相比,该集合提供了更广泛的转录因子覆盖范围,并且在识别受干扰的转录因子方面表现更优。与DoRothEA类似,相互作用根据其调节模式(激活或抑制)进行加权。转录因子合集可以通过decoupleR函数get_collectri进行获取,其中参数split_complexes 用于保留复合物或将其拆分为亚基,默认情况下,建议保留复合物。
tf_net <- decoupleR::get_collectri(organism = 'mouse', #物种human or mouse
split_complexes = FALSE)Activity inference with Univariate Linear Model (ULM):使用ulm推断转录因子(TF)的富集分数。对于数据集中的每个样本(cell)以及网络(tf_net)中的每个转录因子,它拟合一个线性模型,该模型仅根据TF的TF-基因相互作用权重来预测观察到的基因表达。拟合完成后,所获得的斜率的t值即为分数。如果该值为正,则解释为转录因子处于活跃状态;如果为负,则解释为处于不活跃状态。分析的输入与前面通路活性推断类似,需要一个输入矩阵(mat)、一个输入的先验知识网络(net)以及我们想要使用的net中的列名。
tf_acts <- decoupleR::run_ulm(mat = mat,
net = tf_net,
.source = 'source',
.target = 'target',
.mor='mor',
minsize = 5)可视化与通路活性可视化一样:
# Extract ulm and store it in tfsulm in pbmc
sce_cca[['tfsulm']] <- tf_acts %>%
tidyr::pivot_wider(id_cols = 'source',
names_from = 'condition',
values_from = 'score') %>%
tibble::column_to_rownames('source') %>%
Seurat::CreateAssayObject(.)
# Change assay
DefaultAssay(object = sce_cca) <- "tfsulm"
# Scale the data
sce_cca <- Seurat::ScaleData(sce_cca)sce_cca@assays$tfsulm@data <- sce_cca@assays$tfsulm@scale.datacolors <- rev(RColorBrewer::brewer.pal(n = 11, name = "RdBu")[c(2, 10)])
FeaturePlot(sce_cca, features = c("Pax5")) +
ggplot2::scale_colour_gradient2(low = colors[1], mid = 'white', high = colors[2]) +
ggplot2::ggtitle('Pax5 activity')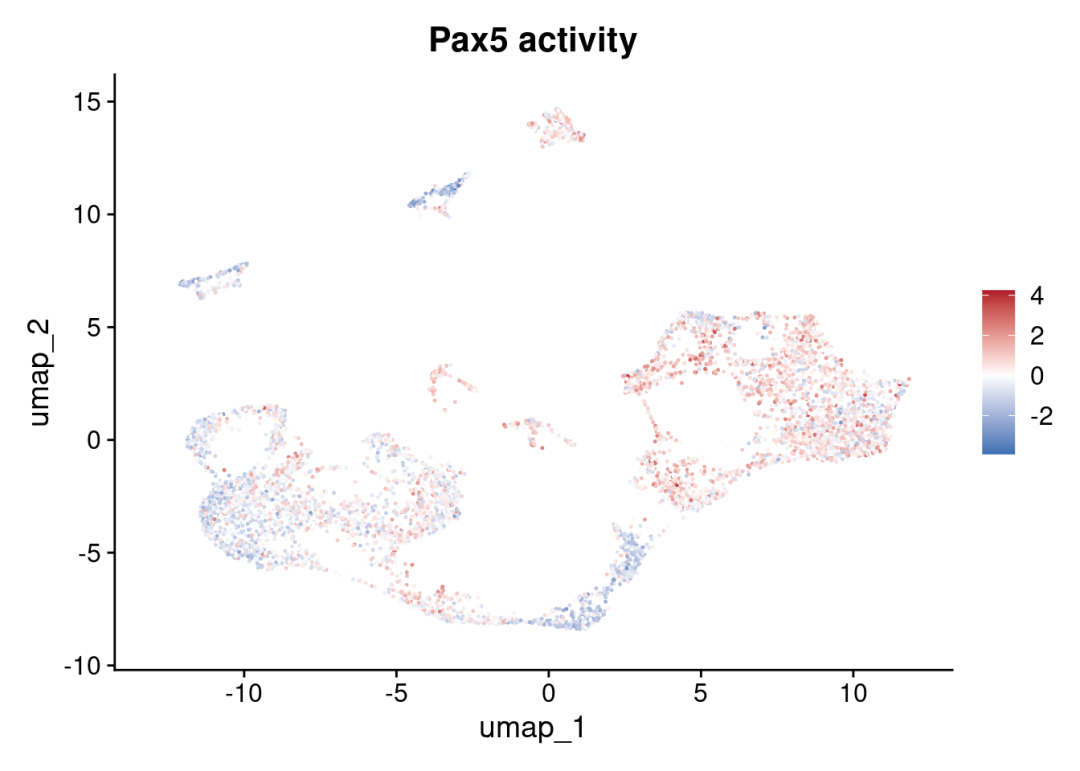
热图可视化每组平均TF活性:展示top25
n_tfs <- 25
# Extract activities from object as a long dataframe
df <- t(as.matrix(sce_cca@assays$tfsulm@data)) %>%
as.data.frame() %>%
dplyr::mutate(cluster = Seurat::Idents(sce_cca)) %>%
tidyr::pivot_longer(cols = -cluster,
names_to = "source",
values_to = "score") %>%
dplyr::group_by(cluster, source) %>%
dplyr::summarise(mean = mean(score))
# Get top tfs with more variable means across clusters
tfs <- df %>%
dplyr::group_by(source) %>%
dplyr::summarise(std = stats::sd(mean)) %>%
dplyr::arrange(-abs(std)) %>%
head(n_tfs) %>%
dplyr::pull(source)
# Subset long data frame to top tfs and transform to wide matrix
top_acts_mat <- df %>%
dplyr::filter(source %in% tfs) %>%
tidyr::pivot_wider(id_cols = 'cluster',
names_from = 'source',
values_from = 'mean') %>%
tibble::column_to_rownames('cluster') %>%
as.matrix()
# Choose color palette
colors <- rev(RColorBrewer::brewer.pal(n = 11, name = "RdBu"))
colors.use <- grDevices::colorRampPalette(colors = colors)(100)
my_breaks <- c(seq(-2, 0, length.out = ceiling(100 / 2) + 1),
seq(0.05, 2, length.out = floor(100 / 2)))
# Plot
pheatmap::pheatmap(mat = top_acts_mat,
color = colors.use,
border_color = "white",
breaks = my_breaks,
cellwidth = 15,
cellheight = 15,
treeheight_row = 20,
treeheight_col = 20) 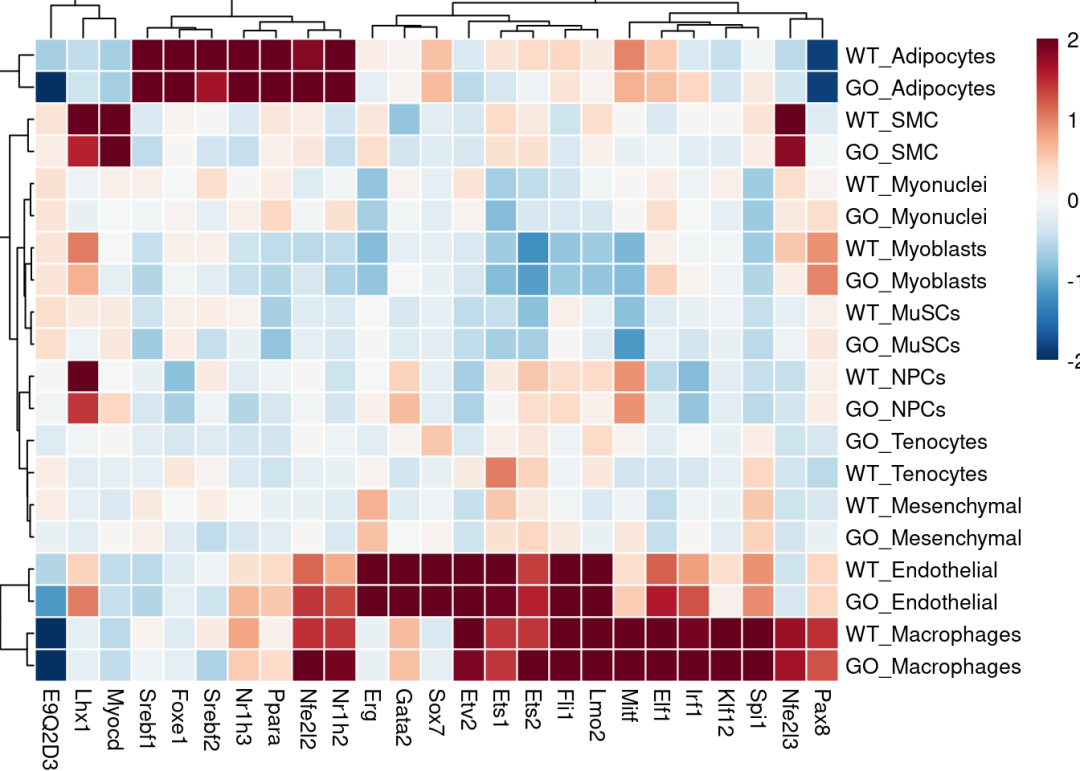
觉得我们分享有用的点个赞再走呗!
本文参与 腾讯云自媒体同步曝光计划,分享自微信公众号。
原始发表:2026-04-24,如有侵权请联系 cloudcommunity@tencent.com 删除
评论
登录后参与评论
推荐阅读
目录

